Featured Article
Technology Spotlight
Please check out our Clinical Diagnostics Assays section for more information or to find manufacturers that sell these products
Emergence of Healthcare-Associated MRSA
Methicillin, the first beta-lactamase-resistant penicillin, was introduced in England in 1959, and strains of methicillin-resistant Staphylococcus aureus (MRSA) were being reported as early as 1961. During the first ten years since those reports, hospital outbreaks of MRSA occurred in Western Europe, Australia, and the United States; and MRSA was eventually recognized to be endemic in hospitals. By 2003, the National Nosocomial Infections Surveillance System estimated that more than 60% of S.aureus isolates in U.S. hospital intensive care units were MRSA.1 Worldwide, similarly high rates have been reported.2
Emergence of Community-Associated MRSA
MRSA infections in children with no healthcare-associated risk factors were reported in the U.S. in the mid-1990s, and there had been scattered case reports of patients with no known healthcare-associated risk factors in the 1980s, including Australian aborigines. The emergence of these so-called community-associated CA-MRSA infections has been very rapid; CA-MRSA infections are now common in most U.S. cities.2,3 Some experts believe that CA-MRSA infections are so prevalent that they have created a public health crisis in emergency departments.2 Worldwide, there have been reports of infections and colonization with CA-MRSA since the early 1990s.3
Features of Healthcare- and Community-Associated MRSA
Studies have shown that there are two epidemiologically and genetically distinct MRSA groups causing invasive MRSA infections: the healthcare-associated (HA)-MRSA strains that evolved and spread in hospitals over the past 40 years, and the CA-MRSA strains that evolved in the community and began to spread about 15 years ago.4 Methicillin resistance is mediated by the mecA gene located in the staphylococcal chromosome cassette mec (SCCmec), a mobile genetic element that may also contain other genetic structures that encode resistance to nonbeta-lactam antibiotics. The HA-MRSA strains carry SCCmec types I through III, and types II and III are large genetic elements that account for the multiresistant phenotype of HA-MRSA. The CA-MRSA strains carry SCCmec IV and V. SCCmec IV and V are small, only carrying resistance to beta-lactam antibiotics; therefore, CA-MRSA strains tend to be susceptible to narrow-spectrum nonbeta-lactams.4,5 In addition, many CA-MRSA strains carry genes for Panton-Valentine leukocidin (PVL), an exotoxin that appears to be partly or wholly responsible for the enhanced pathogenicity of some strains, including the development of necrotic skin lesions. PVL genes are not common in HA-MRSA strains.2,4
Therefore, the HA-MRSA strains are highly resistant to antimicrobials and typically infect persons with healthcare-associated risk factors, including hospitalization, chronic dialysis, antibiotic treatment, and exposure to invasive devices or procedures. These strains tend to cause pneumonia, bacteremia, and invasive infections. CA-MRSA strains are less resistant to antimicrobials and occur in patients without prior healthcare exposure, who are usually previously healthy, younger patients. These strains most commonly cause skin and soft tissue infections, but are also associated with cases of necrotizing pneumonia and severe sepsis.
Status of MRSA Infection Control
Although there is controversy because of differing study methodologies and case definitions for the types of MRSA infections, it is currently thought that rates of HA-MRSA infections have been declining, and that infections with CA-MRSA have increased or remained steady.3,4,7 The epidemiology has been further complicated, because since the early 2000s, HA-MRSA isolates have been found circulating in the community. And, increasingly, MRSA clones bearing SCCmec type IV, particularly the pulsed-field gel electrophoresis DNA fragment USA300, which is the predominant CA-MRSA genotype in the United States, have become common nosocomial pathogens.2,6
The reduction in disease caused by healthcare strains may be a result of increased awareness among hospitals regarding MRSA control;4 however, there have been conflicting results from studies on the efficacy of hospital interventions, which have included implementation of universal nasal surveillance of patients (screening), contact precautions, and hand hygiene.8,9 The varied results may also be a reflection of different study designs. It is noteworthy that MRSA infections have been rare, even in healthcare settings, in Finland, Norway, Sweden, The Netherlands, and Denmark, all of which have had strict surveillance programs for decades.2 By 2011 in the United States, at least 27 states had legislation regarding hospital-acquired infections (HAIs), including MRSA.10 Most states mandate the reporting of HAIs; some also require the implementation of infection control plans, and a few also require universal or high-risk screening.
MRSA Guidelines for Screening
The 2010 screening guidelines11 of the Centers for Disease Control and Prevention (CDC) refer to recommendations from the Clinical and Laboratory Standards Institute, which are as follows: using the cefoxitin disk screen test; the latex agglutination test for penicillin binding protein 2a (PBP2a), which accounts for the resistance to beta-lactam antiobiotics; or a plate containing 6 μg/mL of oxacillin in Mueller-Hinton Agar supplemented with NaCl (4% w/v; 0.68 mol/L). Oxacillin and cefoxitin are used instead of methicillin, which is no longer commercially available in the United States; they also have technical advantages over methicillin. The term “MRSA” is still being used because of methicillin’s historic role. Additional screening tests include nucleic acid amplification tests, such as the polymerase chain reaction (PCR), which are used to detect the mecA gene.
Using the Nasal Swab for MRSA
Nasal screening is the most common approach for MRSA surveillance. However, a study of patients admitted to specialty wards at two large hospitals that looked at screening of various body sites found that although nasal screening performed better than throat, axillary, or perineal screening, it identified only 66% of the true MRSA positives. The authors estimated that realistic screening compliance rates would indicate that nasal swabbing by itself probably detects just over half of true colonization.12 Another author has reported that screening using pooled nose, throat, and perineal swabs cultured on an MRSA-selective chromogenic agar, followed by testing presumptive MRSA colonies using the latex agglutination test for PBP2a, provided positive results after 24 hr of incubation in greater than 95% of true-positive cases.13
MRSA Culture
The following companies offer selective, chromogenic media for the qualitative detection of nasal colonization by MRSA. All products are intended to be used for swab specimens from the anterior nares and are not for diagnosis or to monitor treatment of MRSA infections. All media are incubated at 35–37 °C.
- BD Diagnostic Systems (Sparks, MD) offers BBL™ CHROMagar® MRSA II, a modification of the original BBL CHROMagar MRSA. The selective and differential medium incorporates cefoxitin for identification of MRSA. The plates are incubated for 20–26 hr; MRSA strains produce mauve colonies, which are a result of hydrolysis of the chromogenic substrate. The product is available in packages of 20 plates or cartons of 100 plates.
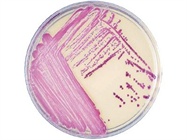
- Bio-Rad Laboratories, Inc. (Hercules, CA) offers MRSASelect™, a selective chromogenic medium for the identification of MRSA within 18–24 hr of incubation. Culture selectivity is based on an unspecified antibiotic/antifungal mixture, and identification is based on the cleavage of the chromogenic substrate by a specific enzyme of S.aureus. MRSA strains produce small pink colonies on this medium. The product is available in packages of 20 plates (90 mm).
- Hardy Diagnostics (Santa Maria, CA) offers HardyCHROM™ MRSA, a selective and differential medium that incorporates unspecified inhibitory and selective agents; MRSA strains produce pink to magenta colonies within 24 hr of incubation. The product is available in bags of 10 plates (15 × 100 mm). Hardy also makes HardyChrom MRSA contact plates to test environmental surfaces. Each plate has a grid molded onto the bottom of the plate to enable enumeration of microbial colonies. The contact plates should be incubated at 35 °C for 16–24 hr, and, if negative at 24 hr, should be incubated an additional 24 hr. Coagulase testing should be performed on any colonies appearing at 48 hr, before reporting as MRSA.
Assays for Penicillin Binding Protein 2a
The following companies offer immunoassays for PBP2a. None of the assays is intended for diagnosis or to monitor treatment of MRSA infections.
- To detect PBP2a from S. aureus culture isolates, Alere™ Inc. (Orlando, FL) offers the Alere PBP2a Culture Colony Test, which is a rapid immunochromatographic assay that uses monoclonal antibodies to detect PBP2a from S.aureus isolates on a culture plate. The antibodies and a control antibody are immobilized onto a nitrocellulose membrane and combined with a sample pad, a blue conjugate pad, and an absorption pad to form a test strip. Colonies may be tested from Tryptone Soy Agar with 5% sheep blood (TSA with blood), Columbia Agar with 5% sheep blood, and Mueller-Hinton Agar. Other media need validation prior to use with this test. The manufacturer recommends testing fresh cultures at 18–24 hr. Results are available within 6 min.
- Hardy Diagnostics offers the MRSA Latex Test for PBP2' (PBP2a), a rapid latex agglutination assay that detects PBP2a in S. aureus isolates. The test consists of a latex reagent sensitized with monoclonal antibody against PBP2a, plus extraction reagents. Extracts are prepared by boiling a suspension of S.aureus cells under alkaline conditions, followed by neutralization and centrifugation steps. The supernatant is then mixed with the latex reagent on a test card, and visible clumping or agglutination within 3 min indicates the presumptive presence of PBP2a. Colonies may be tested from TSA with blood, Columbia Agar with 5% sheep blood, and Mueller-Hinton Agar. The use of fresh (18–24 hr) cultures (grown at 35 °C) is recommended. The total test time is 15 min, and there are 50 tests/kit.
- Thermo Fisher Scientific (Middletown, VA) offers the Thermo Scientific™ Oxoid™ PBP2' (PBP2A) Latex Agglutination Test Kit for isolates of Staphylococcus, as an aid in identifying MRSA and methicillin-resistant coagulase negative staphylococci. The test consists of a latex reagent sensitized with monoclonal antibody against PBP2a, plus an extraction reagent. There are 50 tests/kit.
The following companies offer rapid real-time (RT)-PCR detection of MRSA DNA directly from nasal swabs. None of the assays is intended for diagnosis or to monitor treatment of MRSA infections.
- BD Diagnostics (La Jolla, CA) offers the BD GeneOhm™ MRSA ACP Assay. The test utilizes fluorogenic target-specific hybridization probes for the detection of amplified MRSA DNA. This assay is featured as an improvement over BD’s previous assay, with a simplified work flow that enables detection of MRSA within 2 hr. There is no culture step. The test requires the BD GeneOhm MRSA Lysis Kit (100 tests/kit) and the Cepheid SmartCycler® II starter system with Dx Software. The assay is available in kits of 48 or 200 tests, and provides the Master Mix, Control DNA, Diluent, and SmartCycler reaction tubes (25 μL).
- Cepheid (Sunnyvale, CA) offers Xpert® MRSA, a rapid, moderately complex test. The swab is inserted into a sample reagent vial, which is vortexed, and the sample is dispensed from the vial into the specimen port of the XpertMRSA cartridge, which is then inserted into the GeneXpert® System. Results are available within 66 min. Xpert MRSA is capable of detecting strains with all SCCmec types found in both HA- and CA-MRSA. It is available as 10 or 120 cartridges, plus reagents.
- Roche Molecular Diagnostics (Pleasanton, CA) offers the LightCycler® MRSA Advanced Test. The test is performed on the LightCycler 2.0 instrument and uses swab extraction and mechanical lysis for specimen preparation followed by RT-PCR, and fluorogenic target-specific hybridization probes for detection of amplified MRSA DNA. Results are available in 2 hr for batches of 1–30 samples.
References
- Klevens, M.R.; Edwards, J.R. et al. Changes in the epidemiology of methicillin-resistant Staphylococcus aureus in intensive care units in US hospitals, 1992–2003. Clin. Infect. Dis. 2006, 42, 389–91.
- David, M.Z.; Daum, R.S. Clin. Microbiol. Rev. July 2010, 616–87.
- http://mrsa-research-center.bsd.uchicago.edu/timeline.html.
- Wallin, T.R.; Hern, G. et al. Community-associated methicillin-resistant Staphylococcus aureus. Emerg. Med. Clin. N. Am. 2008, 26, 431–55.
- Ito, T.; Ma, X. et al. Novel type V staphylococcal cassette chromosome mec driven by a novel cassette chromosome recombinase, ccrC. Antimicrob. Agents Chemother. 2004 July, 2637–51.
- http://emedicine.medscape.com/article/971358-overview.
- http://www.cdc.gov/mrsa/statistics/index.html.
- Jain, R.; Kralovic, S.M. et al. Veterans affairs initiative to prevent methicillin-resistant Staphylococcus aureus infections. N. Engl. J. Med. 2011, 364, 1419–30.
- Huskins, W.C.; Huckabee, C.M. et al. Intervention to reduce transmission of resistant bacteria in intensive care. N. Engl. J. Med. 2011, 364, 1407–18.
- http://www.hospitalinfection.org/legislation.shtml.
- http://www.cdc.gov/mrsa/lab/lab-detection.html.
- Matheson, A.; Christie, P. et al. Nasal swab screening for methicillin-resistant Staphylococcus aureus—how well does it perform? A cross-sectional study. Infect. Control Hosp. Epidemiol. 2012, 33, 803–8.
- French, G.L. Methods for screening for methicillin-resistant Staphylococcus aureus carriage. Clin. Microbiol. Infect. 2009Suppl 7, 10–16.
Please check out our Clinical Diagnostic Assays section for more information or to find manufacturers that sell these products