Featured Article

Immunoassays have been the diagnostic of choice for measuring the concentration of molecules in biological fluids for nearly 60 years. Yet, the process used to create a solid-phase immunoassay has changed little—antibody is adsorbed onto a microplate well, microbead or fluidic channel in much the same manner: loosely bound or unbound excess antibody is rinsed away until a stable coating is obtained. This works reasonably well in most applications in which a single antibody is to be coated. However, for multiplexing—in which numerous antibodies need to be immobilized discreetly from one another (e.g., antibody microarray)—passive adsorption has limitations. This is because various antibodies may adsorb onto a particular surface in different ways, depending on their physical-chemical properties; this leads to variations in surface morphology, packing density, surface denaturation and orientation to the surface that may limit availability for antigen binding.
Matson and Little1 found that immunosorbents prepared by linkage of antibodies through oriented coupling to the surface significantly improved antigen-binding capacity compared to antibody immobilized via random coupling. Butler et al.2 showed that less than 3% of monoclonal capture antibody passively adsorbed onto polystyrene was capable of binding antigen.
Others have confirmed that the direct, stochastic immobilization of an antibody to a surface may result in as little as 5–10% of residual binding efficiency for antigen.3 It makes sense that creating antibody microarrays based on oriented immobilization would improve sensitivity. However, from a manufacturing standpoint, other factors come into play, i.e., cost, scalability and robustness. Moreover, there is renewed interest in microfluidic or other chip-based immunosensors for multiplexing in point-of-use or point-of-care applications, especially any that may incorporate mobile connectivity. The ability to migrate developed immunoassays onto new platforms would be desirable.
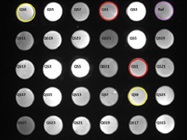
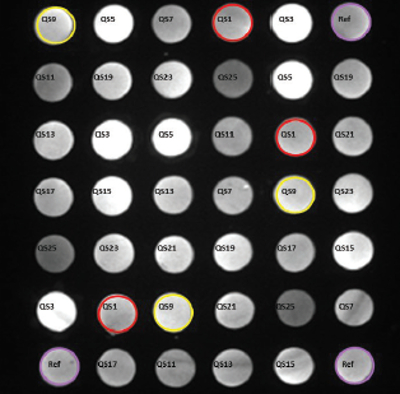 Figure 1 – Location and assignment of the capture oligonucleotides in an A2 Plate well. The covalently linked oligos are printed in triplicate. QS#’s correspond to spots mapped to the complementary oligo that is used for conjugation. Fluorescent spots shown were developed using a pool of biotinylated complementary oligos.
Figure 1 – Location and assignment of the capture oligonucleotides in an A2 Plate well. The covalently linked oligos are printed in triplicate. QS#’s correspond to spots mapped to the complementary oligo that is used for conjugation. Fluorescent spots shown were developed using a pool of biotinylated complementary oligos.Oligonucleotide tethering (QuantiScientifics, Orange, Calif.) offers an unconventional approach to antibody immobilization. Using A2 technology originally developed at Beckman Coulter Inc. (Brea, Calif.), the process is based on the use of microarrays comprising uniquely designed capture oligonucleotides covalently linked to the wells of a 96-well polypropylene microplate (A2 Plate). The microarray in each well is composed of 42 spots in a 7 × 6 pattern of 13 unique oligos represented in triplicate, together with a reference oligo for imaging (Figure 1). Complementary oligonucleotides containing terminal succinimidyl 4-(N-maleimidomethyl)cyclohexane- 1-carboxylate (SMCC) reactive groups are coupled to thiol-modified antibodies in a solid-phase process based on hydrophobic interaction. The reaction and subsequent purification take place in small spin columns (Figure 2a). Once prepared, the oligo-antibody conjugates are mixed together in a hybridization cocktail and incubated with the plate-bound capture oligo array for 1 hour, resulting in the formation of an oligo-tethered antibody array (Figure 2b). The separate oligo-antibody conjugates can be stored as reagents for future use. Because immobilization is thermodynamically driven and based on the sequence-specific, self-assembly of the capture oligo/oligo-antibody conjugate pairs, it is possible to create different antibody arrays in individual wells of the same plate.
There are other advantages to this approach. Reverse hybridization may limit hybrid formation that, in turn, controls the antibody density on spots. Refinement of the oligo pair design and hybridization buffer permits the development of microarrays with uniformity in antibody density and surface morphology. Finally, each capture oligo is extended from the surface via a hydrophilic polymer linker. This may help orient the antibody away from the surface, allowing antigen to be sequestered from the bulk solution.
 Figure 2 – a) Flow chart for oligo-antibody conjugation. b) Process for creating oligo-tethered antibody microarray.
Figure 2 – a) Flow chart for oligo-antibody conjugation. b) Process for creating oligo-tethered antibody microarray.Preparing oligo-antibody conjugates
Conjugates are prepared by solid-phase coupling of purified antibody with an activated oligonucleotide. The antibody is applied to a hydrophobic interaction resin (butyl sepharose) held within the small spin column, where it is adsorbed. A packed-bed volume of 100 µL of the resin is sufficient to bind approximately 100 µg of antibody. With spin columns, the user can prepare several conjugates in parallel using a microcentrifuge. Once bound to the resin, the antibody is modified using a thiolating reagent (iminothiolane) to create free sulfhydryl groups from available lysine residues of the protein. The thiolated antibody is then coupled to an activated form of the desired complementary oligo. This oligo contains a terminal activated ester (SMCC, 4-[N-maleimidiomethyl] cyclohexane-1-carboxylic acid N-hydroxysuccinimidyl ester) that reacts with free –SH groups on the antibody. The entire process takes place using the spin column. At the end of the reaction, the reactants are rinsed from the resin and the purified oligo-antibody conjugate are eluted from the column, ready for use.
Yield
Yields vary, depending on antibody source. Mouse monoclonal antibodies generally produce yields of 40–70%, while the conjugates of polyclonal antibodies from goat or rabbit are prepared with higher yields. An oligo-antibody conjugate obtained from a spin column with a 60% yield will provide approximately 60 µg of the conjugate. Loading a 96-well plate at 1 µg/mL of the conjugate will consume about 5–6 µg, depending on well volume. As a result, 10–12 plates may be prepared from a single conjugation at that loading.
Loading
Each spot of the array should be fully saturated with capture antibody to achieve the highest assay sensitivity and lowest cost per analyte. Simple load isotherms (titer) may be developed from serial dilution of the oligo-antibody conjugate (load) in hybridization buffer. Once hybridized, the series may be analyzed using a fixed concentration of labeled anti-species antibody to determine bound conjugate. For example, if the capture antibody is a mouse monoclonal, a biotinylated goat anti-mouse antibody can be used to determine saturation. A plot of log (signal intensity) versus log (dilution) will result in noticeable inflection at suboptimal loading.
Examples
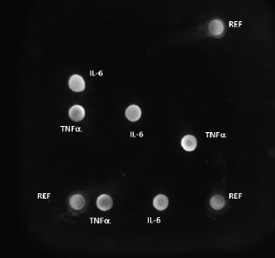 Figure 3 – A2 Plate single-well image showing fluorescent signal for cytokine analytes
Figure 3 – A2 Plate single-well image showing fluorescent signal for cytokine analytesThe analysis of cytokines by multiplex immunoassay is perhaps the most often-cited application. Rossi’s group4 used the A2 platform in their examination of cytokine expression during induction of human CD34+ progenitor cells by siRNA. Arrieta et al.5 evaluated levels of cytokines in plasma and urine in an effort to differentiate sepsis from urinary tract infections in infants. They found the urine/plasma ratio for IL-6 to be a good indicator for screening infants under 3 months of age.
Figure 3 shows a single well from which cytokines IL-6 and TNFα will be analyzed using the A2 microarray reader. The system serves as a controller for automatically reading the A2 Plate wells and provides data analysis and reporting (Figure 4). Data can be exported in Excel (Microsoft, Inc., Redmond, Wash.) files.
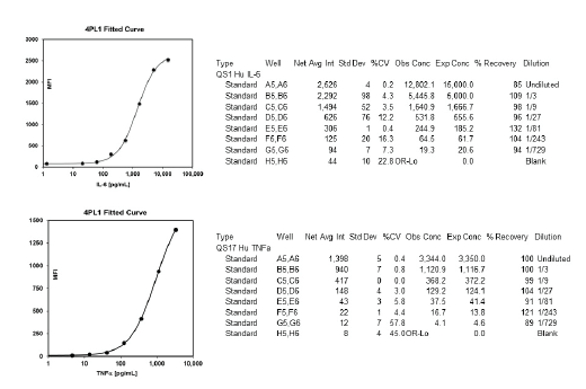 Figure 4 – Reports for cytokines analyzed using the A2 multiplex immunoassay system.
Figure 4 – Reports for cytokines analyzed using the A2 multiplex immunoassay system.Migration to other surfaces
The A2 oligo-tethering format has also been used on polypropylene films and other thermoplastics. The more common planar substrates for microarrays are glass slides and polystyrene plates. Covalent attachment of amine-terminal capture oligos to glass slides is straightforward using slides derivatized with epoxy silane coatings. The A2 multiplex immunoassay slide contains eight subarrays in a 2 × 4 well pattern of the capture oligo array. Correlation to the A2 Plate has been verified, e.g., IL-6 (r2 = 0.97) (see www.quantiscientifics.com).
The attachment of short strands of DNA to polystyrene is more difficult, requiring either photo cross-linking or preparation of chemically activated polymer. A bridging molecule that has a high affinity for the substrate can also be employed.
Immobilization of A2 oligonucleotides onto polystyrene was accomplished via indirect attachment using proteins such as bovine serum albumin (BSA). Tronic6 calculated the surface concentration of BSA at approximately 400 ng/cm2 for a polystyrene substrate but less than 100 ng/cm2 for glass. Oligo-BSA conjugates were prepared and printed onto high-bind polystyrene strip well plates (Greiner Bio-One, Monroe, N.C.) (see Figure 5).
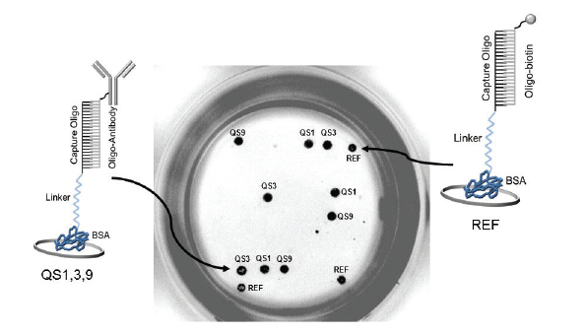 Figure 5 – Fluorescent imaging of a single polystyrene strip well with capture oligos immobilized using BSA as a bridge to the substrate surface
Figure 5 – Fluorescent imaging of a single polystyrene strip well with capture oligos immobilized using BSA as a bridge to the substrate surface.
BSA conjugates (QS1, 3 and 9 and Ref; see Figure 5) were printed in triplicate in polystyrene plate wells. While minor changes in the hybridization buffer and blocking conditions were required, development of the immunoassay is similar to that based on use of the A2 Plate. Good correlation was shown (r2 = 0.9965) when QS3-anti-human FGF (fibroblast growth factor)-acidic analyzed by fluorescent ELISA using the A2 polypropylene plate was compared with QS3-BSA anti-human FGF acidic colorimetric ELISA using the strip well polystyrene plate.
The A2 OnePlex strip well plate is made of oligo-BSA spotted in a 3 × 3 array pattern. Complementary oligo-antibody conjugate was applied to create a single-plex immunoassay plate. This can be developed and the signal read by imaging fluorescent spots using a microarray imaging reader or by applying a colorimetric reagent (solution phase) for use with a conventional microplate reader. Correlation (r2 = 0.9673) between the two formats for a singlet ELISA for IL-6 is shown in Figure 6. Arraying offers an economical, rapid and controllable coating process with flexibility for on-demand applications. Individual strips or wells can be prepared with the same or different capture antibodies as required using oligo-tethering.
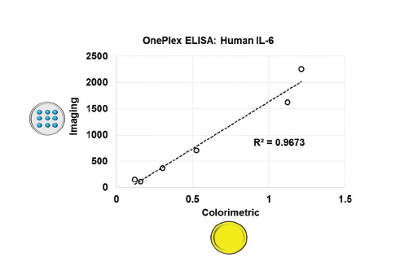 Figure 6 – Correlation between IL-6 ELISAs developed in strip wells using fluorescent imaging or colorimetric assays.
Figure 6 – Correlation between IL-6 ELISAs developed in strip wells using fluorescent imaging or colorimetric assays.Summary
Oligonucleotide tethering of antibodies provides a flexible means with which to develop antibody arrays for quantitative multiplex immunoassay applications. Conjugation of capture oligonucleotides with proteins or other biopolymers that strongly adsorb to polystyrene (or other substrates) enables migration of the technology for use on new platforms and devices.
References
- Matson, R.S. and Little, M.C. Strategy for the immobilization of monoclonal antibodies on solid-phase supports. J. Chromatogr.1988, 458, 67–77.
- Butler, J.E.; Nessler, L.N. et al. The physical and functional behavior of capture antibodies adsorbed on polystyrene. J. Immunol. Meth.1992, 150, 77–90.
- Schramm, W. and Paek, S.-H. Antibody-antigen complex formation with immobilized immunoglobulins. Anal. Biochem.1992, 205(1), 47–56.
- Robbins, M.A.; Li, M. et al. Stable expression of shRNAs in human CD34+ progenitor cells can avoid induction of interferon responses to siRNAs in vitro. Nat. Biotechnol.2006, 24, 566–71.
- Arrieta, A.; Osborne, S. et al. Pilot Study Evaluating Urine Plasma Ratios of Inflammatory Cytokines to Allow for Discrimination of Urinary Tract as the Source of Serious Bacterial Infections in Young Febrile Infants. Poster #241, Infectious Diseases Society of America, San Diego, Calif., 2015.
- Tronic, E. Ph.D. dissertation. Surface analysis of adsorbed proteins: a multi-technique approach to characterize surface structure. U. Washington, pub. no. 3552866; http://search.proquest.com/ docview/1313207756
Robert S. Matson, Ph.D., FACB, is president, QuantiScientifics LLC, 1920 E. Katella Ave., Ste. S, Orange, Calif. 92867, U.S.A.; tel.: 714-624-0810; e-mail: [email protected] ; www.quantiscientifics.com