Featured Article

Automation and algorithms accelerate the use of gigantic datasets
More data and opportunities—as well as expectations—make up today’s focus of analyzing images from microscopy. Not long ago, researchers used imaging data in figures for papers to confirm findings, not so much to quantify them. “Presently, it is rare for a reviewer to allow an image to appear in a manuscript without some sort of quantitative analysis,” says Douglas Richardson, director of the Harvard Center for Biological Imaging (Cambridge, MA). “As all experiments are unique, there is not a one-size-fits-all analysis.” That means that scientists must often customize solutions, which requires skill in computer science and developing algorithms.
Getting the most from analyzing images depends on every step. According to Gregg Kleinberg, vice president new business development at Midwest Information Systems (Villa Park, IL), “The most significant challenge for users is the issue of sample preparation.” He adds, “Since all image analysis, including our PAX-it! Image Analysis software, requires either grayscale or RGB—red, green, blue—thresholding, there must be sufficient contrast in order to separate the objects of interest from the background and from other objects within the field of view.” As an example, he points out the need to polish a metallic sample and chemically etch it to reveal grain boundaries sufficient for automatic image analysis. As he explains, “I recently experienced this with a customer who was trying to make automatic cell measurements in a black polystyrene material, where you could see the cells in a scanning electron microscope image, but the background colors were the same as the cells.” To address that challenge, Kleinberg and his colleagues developed a new method of sample preparation.
So, getting the best analysis begins with sample processing, and then involves every step.
Adding automation
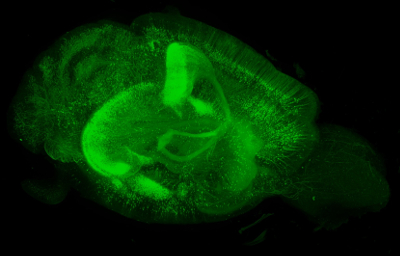 In a genetically modified mouse brain expressing enhanced green fluorescent protein under the Thy-1 promoter, more than 70,000 2-S images were acquired by lightsheet microscopy to produce this 3-D image, which represents over 500 gigabytes of data. (Image courtesy of Douglas Richardson.)
In a genetically modified mouse brain expressing enhanced green fluorescent protein under the Thy-1 promoter, more than 70,000 2-S images were acquired by lightsheet microscopy to produce this 3-D image, which represents over 500 gigabytes of data. (Image courtesy of Douglas Richardson.)Today’s image-acquisition systems use automation to collect more data in far less time. “Experiments that previously struggled to attain an n of 3 are now easily automated to image hundreds or even thousands of samples,” Richardson notes. “This produces vast amounts of data—terabytes—that are difficult to store and cannot be manually analyzed.” He adds, “Only a fully automated computational pipeline can accomplish this.”
The amounts of microscopy data, though, are good and bad—good in providing more detail, but challenging to handle at times. From Duke University’s (Durham, NC) Light Microscopy Core Facility, director Lisa Cameron says, “The biggest issue is handling large files and being able to visualize 3-D renderings, especially multichannel over time.” Light sheet fluorescence microscopy (LSFM), for example, generates so many gigabytes of data so fast that transferring and viewing the files gets tricky. “This is our bottleneck now,” she says.
With so much more microscopy data to analyze, scientists need the tools for the task. Graphic processing units (GPUs) improve computational options. “Although it cannot be applied in every situation,” Richardson explains, “for image analysis methods that can be parallelized, GPU processing can facilitate image analysis on a high-end workstation that in the past needed an excessive amount of processing time or access to a cluster computer.”
In addition to improved computer hardware, advances in algorithms matter, too. “Machine learning and deep learning are having an impact in image classification and segmentation,” Richardson says. “If you are able to produce a high-quality training data set, these techniques can provide accurate image classifications.”
Plus, some of the world’s top computing capabilities now live in the cloud. “This provides everyone access to high-level computing power that in the past was restricted to only top-level institutions,” Richardson says.
Simpler seeing
 Automated imaging systems simplify data acquisition and analysis of microscopy samples. (Image courtesy of Molecular Devices.)
Automated imaging systems simplify data acquisition and analysis of microscopy samples. (Image courtesy of Molecular Devices.)The challenges of managing so much imaging data and analytical options create a niche for complete systems. “It can be difficult to maintain consistent, reliable results with the precise nature of acquiring images and distilling into numerical data,” says Grischa Chandy, manager product marketing and applications, cellular analysis at Molecular Devices (Sunnyvale, CA). “This is where automated imagers, like the ImageXpress Nano system, can provide a robust, reliable platform for acquiring and processing images using slides or multiwell plate formats.”
This system also provides integrated image-analysis capability. As Chandy notes, it comes with “25 predefined analysis protocols that can get users started quickly, while still having the flexibility to create customized protocols.” The protocols provide measurements ranging from cell counts to characterization of neurite outgrowth. Moreover, this system accommodates label-free and fluorescent assays.
The ImageXpress Nano uses browser-based software, which simplifies data sharing and collaborative image analysis. “Being tablet friendly, the software is touchscreen compatible, providing exceptional usability and allowing precise control,” Chandy explains. “The software includes many visual tools for optimization of acquisition and analysis that makes it easy to use, even for novice users.”
Amazing outcomes
Despite some of the ongoing challenges of analyzing images from microscopy, the capabilities would seem amazing by even the standards of five years ago. Putting all of the power together opens new opportunities. As one example, Richardson describes rendering a 5-terabyte dataset in three dimensions on a laptop and sharing it with colleagues. “Another interesting advance is the ability to track individual cells in a live sample over time,” he says. “This has been done for years on relatively small samples in 2-D, but now, every cell in complex multi-cellular organisms can be followed over long periods of time in 3-D.”
Scientists can also compare datasets to each other or to reference atlases. As Richardson says, this “allows for the detection of phenotypic differences between control and disease models in 3-D.”
Some approaches to imaging look very futuristic. “Now, using PAX-it! Image Analysis software and motorized components for microscope retrofit, you can load multiple samples into a fixture on the motorized microscope stage, and using software with scripting, you can automate a stage move to a programmed location, trigger an autofocus, trigger an image capture, launch analysis of that image and add the data to a cumulative report, which concludes and displays for review, once the samples finish running,” Kleinberg explains.
As the tools for analyzing images from microscopy continue to improve, even more applications will arise. Even now, this technology impacts fields from clinical science and forensics to life sciences and semiconductors. But even as some tools get simpler to use, research scientists should expect to improve their mathematical and computational skills to develop methods on the edge of what is possible. Only then can we keep improving our understanding of the microscopic things in our world.
Mike May is a freelance writer and editor living in Texas. He can be reached at [email protected].