Featured Article
The practice of clinical laboratory medicine is dependent on the discovery and availability of new and applicable technological tools, and clinical laboratories worldwide have been the beneficiaries of a remarkable number of technological advances. Consequently, the practice of laboratory medicine has been transformed from a secondary to a central role in healthcare delivery.1 Specific arenas in which technological advancements have occurred include analytical systems, analytical methodology, numerous relevant technologies, and molecular diagnostics.
Analytical systems
Analytical systems for performing clinical assays are classified as either continuous flow analyzers or discrete analyzers.
Continuous flow analyzers
In this type of analyzer, an aliquot of the sample is introduced into a tube of continuous flowing reagent(s) followed by mixing and sequential passage through a number of processing modules ending in a flow-through monitoring sensor (Figure 1).2
 Figure 1 – Schematic of continuous flow analyzer with two examples.
Figure 1 – Schematic of continuous flow analyzer with two examples.Discrete analyzers
With a discrete analyzer, aliquots of sample and reagent(s) are introduced into, mixed, and processed within a discrete chamber (often a cuvette) (Figure 2). Thus, each sample and its ensuing chemical reaction has its own physical and chemical space and maintains it as each chamber then travels down a prescribed pathway in either a linear or circular fashion.3
 Figure 2 – Schematic of a discrete analyzer with an example of an automated discrete system.
Figure 2 – Schematic of a discrete analyzer with an example of an automated discrete system.In practice, discrete analyzers have less carryover and are faster and easier to calibrate, operate, and automate.4,5
Automated analyzers
In current practice, a variety of automated devices are found in clinical laboratories that have been designed to improve the quality, throughput, and efficiency of laboratory testing.6 They vary from automated instruments to completely automated systems termed total laboratory automaton (TLA). These are found in high-volume laboratories that produce results for a wide variety of analytes (Figure 3).
 Figure 3 – Example of a totally automated analytical system.
Figure 3 – Example of a totally automated analytical system. Key developments that have led to TLA include robotic transport systems, robotic pipetting systems, bar-coding devices, a wide variety of measurement devices, and powerful computer systems with their prerequisite software.
Point-of-care analyzers
Point-of-care testing (POCT) is a mode of testing in which the analysis is performed close to the patient, for example, in the home and in primary, secondary, and tertiary healthcare facilities.7 Such testing has multiple advantages, including reduced turnaround time, operational simplicity, reduced cost, and opportunity for timely and informed discussion with the patient.
The use of POCT for healthcare has increased during the last five decades since its introduction, principally driven by technology developments and changes in healthcare delivery that are aimed at delivering less costly and more effective care closer to the patient’s home.7 Clinically, the first use of POCT was the use of reagent-loaded paper strips for measuring blood glucose concentrations in diabetics.5
Recently, the analytical scope of POCT devices has increased as a result of integration of advances made in miniaturization, lateral flow microfluidics, sensors, bar coding, informatics, and reagent and materials manufacturing.
Relevant technologies
Technologies that have played a significant role in the development of the analytical devices discussed above include:
Robots
The Robot Institute of America defines a robot as: “A reprogrammable, multifunctional manipulator designed to move materials, parts, tools, or specialized devices through various programmed motions for the performance of a variety of tasks.” Robots are widely employed in today’s automated systems.
Microfabrication
Microfabrication is the process of fabrication of miniature structures of micrometer scales and smaller. Microelectromechanical systems (MEMS) is the technology of very small devices. Examples include miniaturization of POC analyzers, mobile phones, integrated circuits, and sensors.
Microfluidics
Microfluidics is the science and technology of manipulating and controlling fluids, usually in the range of microliters (10-6) to picoliters (10-12), in networks of channels with lowest dimensions from tens to hundreds of micrometers. It is an important technology that is used in the development of POC devices.
Nanotechnology
Nanotechnology is manipulation of matter on an atomic, molecular, and supramolecular scale. It is a technology that shows great promise in the development of biomarker discovery, cancer diagnostics, and detection of infectious microorganisms.8 Categories of nanodiagnostic technologies available include:
- Biosensors
- Biochips
- Nucleic acid technologies
- Bio-barcode assays
- DNA nanomachines
- Nanoparticle biolabels
- Microarrays
- Proteomic technologies
- Nanopore technology
- Nano-immunoassays.
Computer technology
The COLOSSUS and ENIAC were vacuum tube computers first developed by the British and United States governments. Later generations of computers were fabricated using integrated circuits of transistors with each generation being faster, having increased memory, and operated with more powerful and applicable software.
Information technology
Information technology (IT) is the use of computers to store, retrieve, transmit, and manipulate data. IT has revolutionized the means by which a modern clinical laboratory generates, communicates, manages, and utilizes relevant information. Processes used for this purpose include communication devices such as e-mail, texting, search engines, electronic books, electronic journals, smart phones, and networks.
Analytical methodology
Considerable progress and innovation have been made in the introduction of advanced analytical methodologies that complement the analytical systems discussed above. Significant examples include enzyme immunoassay and molecular diagnostics.
Enzyme immunoassay
Immunoassay is a versatile and useful analytical technique that utilizes antibodies as reagents to measure an antigen of interest.9 They were first introduced into clinical laboratories with the invention in the late 1950s by Yalow and Berson of a radioimmune assay (RIA) for measuring insulin.10
RIA methods were once popular, but their use has decreased because of safety concerns and lack of sensitivity. These problems were avoided with the invention of enzyme immunoassays (EIAs) that replaced radioisotopes with enzymes as labels.11
Different types of EIA include:
- Enzyme-linked immunosorbent assay (ELISA)
- Enzyme multiplied immunoassay (EMIT)
- Cloned enzyme donor immunoassay (CEDIA).
These assays initially used photometric labeling and measurement; however, luminescence labels are now used because of their increased sensitivity.9 In practice, due to their increased analytical sensitivity and adaptability to be added to the assay menu of automated instruments and systems, enzyme immunoassays have increased the scope of such systems and they are now widely used for measuring proteins, hormones, and drugs.
Molecular diagnostics
The discovery of the double helix as DNA’s basic structure and the success in determining the sequence of nucleotide base pairs in the human genome were fundamental findings that led to the development of several molecular diagnostic techniques. These are now used by clinical laboratories to analyze biological markers in the genome and proteome and how cells express their genes as proteins (Figure 4).12 This information is being used to diagnose and monitor disease, detect risk, and select appropriate therapies.
 Figure 4 – Molecular diagnostics (MDx)—a technique used to analyze biological markers in the genome and proteome.
Figure 4 – Molecular diagnostics (MDx)—a technique used to analyze biological markers in the genome and proteome.Categories include:13,14
Amplification techniques
Molecular diagnostic methods often require an amplification step to detect very low concentrations of nucleic acids in a background of complex genomic structure.12 Techniques used for this purpose include those that increase the 1) amount of the target nucleic acid, 2) intensity of the detection signal, or 3) amount of probe.
Thepolymerase chain reaction (PCR)is a widely used amplificationtechnique based on increasing the amount of target nucleic acid. In practice, it is used to amplify selected sections of DNA or RNA for subsequent analysis. This technique can achieve over a million-fold amplification in less than an hour.
Detection techniques
Molecular diagnostic methods use generic and specific methods to detect nucleic acids. In the former, ultraviolet spectrophotometry and fluorometry coupled with fluorescent staining dyes are commonly used to measure nucleic acids. Specific methods utilize sequence-specific primers or probes with fluorescent or electronic detection.
Discrimination techniques
Discrimination techniques include electrophoretic assays that physically separate nucleic acids based on molecular size or shape and hybridization methods. Hybridization methods are based on the ability of single-stranded nucleic acids to form specific double-stranded hybrids (Figure 5). The technique requires a probe and target nucleic acid that, when mixed, allow for a complementary base-pairing after which there is a need to detect any double-stranded nucleic acids. Types of assays include both solid- and solution-phase assays and hybridization microarrays (also known as DNA arrays, DNA chips, and biochips) (Figure 6).
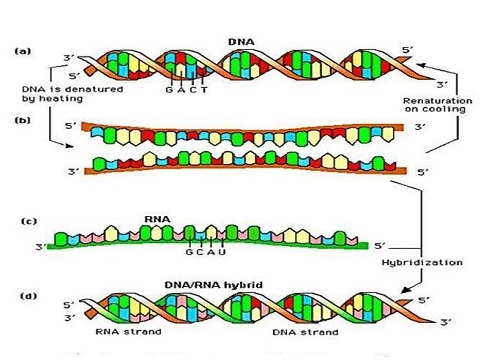 Figure 5 – Nucleic acid hybridization.
Figure 5 – Nucleic acid hybridization. Figure 6 – DNA microarrays—a grid of DNA segments of known sequence that is used to test and map DNA fragments, antibodies, or proteins.
Figure 6 – DNA microarrays—a grid of DNA segments of known sequence that is used to test and map DNA fragments, antibodies, or proteins.Whole genome sequencing
This is the process of determining the complete DNA sequence of an organism’s genome and provides the most comprehensive collection of an individual’s genetic variation.With the birth of the new discipline of personalized medicine, whole genome sequence data is considered by many as an important necessity to guide therapeutic intervention and there has been debate as to should everybody’s genome be determined. With new techniques for sequencing nucleic acids in an efficient, timely, and cost-effective manner, this may be an attainable goal.
In the last five decades, incredible innovations and technological developments have shaped how modern clinical laboratories are equipped, staffed, and operated. In the future it is expected that this trend will continue and will be driven by technological developments in disciplines such as microengineering, advanced robotics, microfabrication, microfluidics, nanotechnology, advanced materials, and particularly molecular diagnostics. It will be fascinating to follow how the clinical laboratory of the future will evolve and how its operation will impact the practices of laboratory medicine and healthcare delivery.
References
- Rifai, N.; Topol, E. et al. Disruptive innovation in laboratory medicine. Clin. Chem. 2015, 61(9), 1129–32.
- Skeggs, L.T. An automatic method for colorimetric analysis. Amer. J. Clin. Path. 1957, 28, 311–22.
- Northam, B.E. Discrete analysis systems. J. Clin. Path. 1969, 22, Suppl. (Coll. Path.), 3, 42–50.
- Rosenfeld, L. Four Centuries of Clinical Chemistry. Taylor & Francis: New York, NY, 1999.
- Berger, D. A brief history of medical diagnosis and the birth of the clinical laboratory. Part 2—laboratory science and professional certification in the 20th century.MLO Aug 1999, 31(8), 32–4, 36, 38.
- Hawker, C.D.; Genzen, J.R. et al. Automation in the Clinical Laboratory. In Rifai, N.; Horvath, A.R., Eds. Tietz Textbook of Clinical Chemistry and Molecular Diagnostics, 6th ed. Elsevier: St. Louis, MO, 2018, p. 370.
- St. John, A. and Price, C.P. Point-of-Care Testing. In Rifai, N.; Horvath, A.R. et al., Eds. Tietz Textbook of Clinical Chemistry and Molecular Diagnostics, 6th ed. Elsevier: St. Louis, MO, 2018, p. 371.
- Jain, K.K. Applications of nanobiotechnology in clinical diagnostics. Clin. Chem. 2007, 53, 1–8.
- Kricka, L. and Park, J.W. Immunochemical Techniques. In Rifai, N.; Horvath, A.R. et al., Eds. Tietz Textbook of Clinical Chemistry and Molecular Diagnostics, 6th ed. Elsevier: St. Louis, MO, 2018, p. 371.
- Yalow, R.S. and Berson, S.A. Immunoassay of endogenous plasma insulin in man. J. Clin. Invest. 1960, 39, 1157–75.
- Rubenstein, K.E.; Schneider, R.S. et al. “Homogeneous” enzyme innunoassay: new immunochemical technique. Biochem. Biophys. Res. Commun. 1972, 47, 846–51.
- Lo, W.M. and Chiu, R.W.K. Principles of Molecular Biology. In Burtis C.A. and Bruns, D.E., Eds. Tietz Fundamentals of Clinical Chemistry and Molecular Diagnostics, 7th ed. Elsevier: St. Louis, MO, 2015, pp. 899–914.
- Wittwer, C.T. and Makrigiorgos, G.M. Nucleic Acid Techniques. In Rifai, N.; Horvath, A.R. et al., Eds. Tietz Textbook of Clinical Chemistry and Molecular Diagnostics, 6th ed. Elsevier: St. Louis, MO, 2018, pp. 959–94.
- Green, E.D. and Guyer, M.S. Charting a course for genomic medicine from base pairs to bedside. Nature 2011, 470, 204–13.
Carl Burtis is an Oak Ridge National Laboratory emeritus, Oak Ridge, TN 37830, U.S.A.; e-mail: [email protected]