Featured Article

The detection and quantitation of proteins in biological samples is routine in applications from basic protein characterization to clinical diagnostic testing and drug development. Common methods for protein detection include enzyme-linked immunosorbent assay (ELISA), dot blot and Western blot.1 ELISA uses colorimetric or fluorescent detection of antibodies bound to antigen in solution, which has been previously captured by a second antibody or immobilized on a plate surface.2 Dot blot is a simpler variant of other molecular blotting techniques in that, like ELISA, the sample can be a mixture of proteins, as these techniques do not require separation of proteins prior to detection.3 Western blot, or protein immunoblot, is one of the oldest and most widely used methods for protein detection. The technique requires homogenization of biological materials such as cultured cells or tissue extracts, followed by electrophoresis to separate the resultant protein mixture; the rate at which proteins migrate through the gel in response to electrical current is dependent on intrinsic properties of the protein such as molecular weight. After protein is transferred from the separating gel onto a membrane, the membrane is incubated in a solution containing an antibody that binds to the protein of interest, conjugated to or followed by a colorimetric or fluorescent reporter. The resultant bands are compared to a standard for determination of the targeted protein’s molecular weight. First described in 1979, Western blotting is still widely used to determine the presence or absence of a protein in a biological sample.
Quantitative data from large populations, with single-cell precision
A significant limitation of bulk protein detection assays like Western blot is the need to homogenize cells or tissue, which results in a loss of expression information from individual cells in a population. Techniques that detect and report signal from individual cells can provide quantitative data from large populations about the protein target or other marker levels, from cells of varying phenotype, developmental state or health status.
Flow cytometry addresses this need for quantitative data from significant cell populations by interrogating individual cells for the presence and relative strength of signal from fluorescent reagents or antibodies. However, traditional flow cytometers typically require extensive operator training and expertise, and sheath fluid-based systems are characterized by extensive setup and shutdown, as well as considerable cost to purchase, operate, and maintain.
The Muse cell analyzer from MilliporeSigma (Billerica, Mass.) provides researchers with simple, affordable access to the quantitative data that flow cytometry provides for measuring markers of viability, mitochondrial health, protease activity and more. Using microcapillary fluidics and preoptimized reagents, this compact, portable cell analyzer requires minimal setup and no expertise to operate, and provides a rapid, simplified alternative to more time-consuming methods such as Western blot (that may also demand considerable technical expertise) for routine analysis of cell culture health and the effects of compounds in toxicology studies and drug discovery screening (Figure 1).
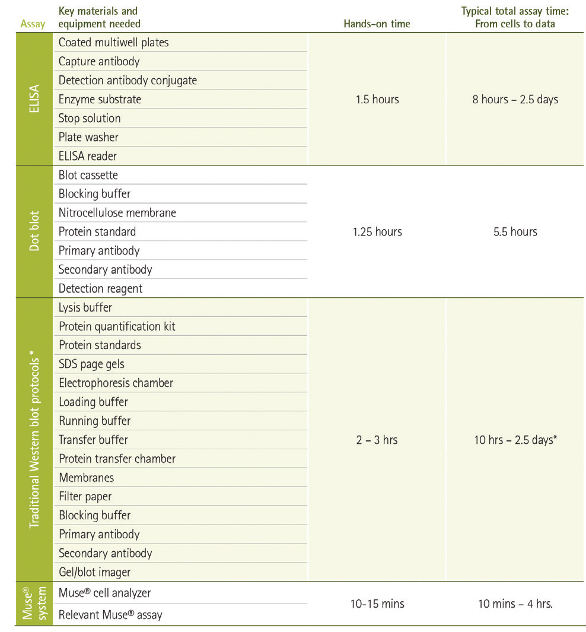 Figure 1 – Workflow comparison among protein detection methods. For proteins not already in solution, bulk methods require the lysis of samples to render proteins accessible to detection. Reagents, supplies and equipment for separation, transfer, probing, detection and imaging may be needed, depending on the technique. Lengthy primary antibody incubations and serial washes following each binding step can result in significant hands-on and total elapsed time for routine protocols. The Muse cell analyzer employs preoptimized reagent cocktails to minimize variation and instrument setup, resulting in substantial reductions in time spent on the bench and total start-to-result time. *During immunodetection, significant time savings can be achieved by use of a vacuum-driven method such as the SNAP i.d. 2.0 system for Western blotting (MilliporeSigma).
Figure 1 – Workflow comparison among protein detection methods. For proteins not already in solution, bulk methods require the lysis of samples to render proteins accessible to detection. Reagents, supplies and equipment for separation, transfer, probing, detection and imaging may be needed, depending on the technique. Lengthy primary antibody incubations and serial washes following each binding step can result in significant hands-on and total elapsed time for routine protocols. The Muse cell analyzer employs preoptimized reagent cocktails to minimize variation and instrument setup, resulting in substantial reductions in time spent on the bench and total start-to-result time. *During immunodetection, significant time savings can be achieved by use of a vacuum-driven method such as the SNAP i.d. 2.0 system for Western blotting (MilliporeSigma).Data may mask population heterogeneity
Data that can be quantified are increasingly important in the life sciences, as quantitative data are objective and therefore considered morereliable and are subject to statistical analysis. Quantitative data are assumed to be more representative of populations than qualitative data, and therefore must be characterized both by significant sample size and by the capacity to measure individual events in a sample.
A constraint of bulk assays is that they can only return data that is qualitative or semiquantitative at best (Figure 2). Although spectrophotometry and densitometry can be used to transform sample well color or the size of a blot or band into numerical values for comparison of relative signal intensity among samples, these methods require homogenization of all the cells or tissue in a particular sample. Western blot also relies on concurrent electrophoresis of a mixture of proteins of known weight to create a standard, or “ladder” of bands on the blot, to which positive bands from sample lanes are compared for confirmation of the protein’s identity (Figure 2a, right panel).
 Figure 2 – Common bulk immunodetection assay results contrasted with quantitative data. a) ELISA, left panel, uses a colorimetric or a fluorescent detection reagent. Spectrophotometry can be used to transform signal intensity into numerical values, but signal is a mean from all cells or cell products in a sample, as is the case with dot blot (middle panel). The right panel is an example of fluorescent detection of immunoblot (Western blot), showing the “ladder,” or molecular standard, in lane 1. b) Left panel, Western blot for validation of anti-H2A.X antibody. Recombinant H2A.X (lane 1), recombinant histone H2A (lane 2) and acid-extracted proteins from HeLa cells (lane 3) were probed with anti-histone H2A.X. The right panel shows representative data from the Muse H2A.X activation dual-detection assay, which uses two directly conjugated antibodies against the unmodified and phosphorylated histone target to map signal from every cell in the sample onto a scatterplot. Absolute numbers and percent of cells activated in the sample are automatically calculated and displayed on the “Statistics” tab.
Figure 2 – Common bulk immunodetection assay results contrasted with quantitative data. a) ELISA, left panel, uses a colorimetric or a fluorescent detection reagent. Spectrophotometry can be used to transform signal intensity into numerical values, but signal is a mean from all cells or cell products in a sample, as is the case with dot blot (middle panel). The right panel is an example of fluorescent detection of immunoblot (Western blot), showing the “ladder,” or molecular standard, in lane 1. b) Left panel, Western blot for validation of anti-H2A.X antibody. Recombinant H2A.X (lane 1), recombinant histone H2A (lane 2) and acid-extracted proteins from HeLa cells (lane 3) were probed with anti-histone H2A.X. The right panel shows representative data from the Muse H2A.X activation dual-detection assay, which uses two directly conjugated antibodies against the unmodified and phosphorylated histone target to map signal from every cell in the sample onto a scatterplot. Absolute numbers and percent of cells activated in the sample are automatically calculated and displayed on the “Statistics” tab.Conclusion
Immunoblot and immunosorbent assays, particularly those that are enzyme-linked, continue to be among the most popular methods for protein detection in the life sciences. They provide clear data confirming the presence, relative abundance or presumed identity of proteins in a cell or tissue sample, and are amenable to measuring virtually any protein for which an epitope binder (such as an antibody) can be developed. These methods are limited, however, in their ability to capture expression information from individual cells in a population. With Western blot, in particular, bulk methods may be prohibitive for routine screening because they are time-consuming and require expertise to optimize reagents and obtain, interpret and troubleshoot results.
Simplified flow cytometry-based analysis presents a rapid, uncomplicated alternative to methods such as immunoblot, especially for routine and frequent screening of cell cultures or for response of cell models to compounds in development for chemotherapeutics, drug discovery, cosmetics and similar applications. The Muse system also incorporates assays for detection using familiar cell status indicators that do not rely on antibody–protein interactions. These include membrane integrity dyes and reagents that fluoresce on nucleic acid intercalation, such as 7-AAD (7-aminoactinomycin D) for viability, and mitochondrial membrane potential indicators. The Muse software automatically provides cell-by-cell results from these reagents, unlike microscopy or other low-throughput, time-consuming or subjective techniques for measuring their signal. Muse assays are selected to provide an efficient means for the most essential viability, cell health and signaling screening, and reagents are preoptimized to minimize variation and the need for complex setup adjustments that characterize traditional open-system cytometers.
Despite its compact size and simplified operation, the Muse delivers the same single-cell data as larger, more costly and complex systems. The availability of rapid cell analysis returning quantitative results without the need for reagents, supply and equipment investment or trained personnel can significantly impact compound screening and cell culture model paradigms in the pharmaceutical, food and beverage and life science industries.
References
- Corley, R.B. A Guide to Methods in the Biomedical Sciences. Springer: New York, N.Y., 2005.
- Lequin, R.M. Enzyme immunoassay (EIA)/enzyme-linked immunosorbent assay (ELISA). Clin. Chem. 2005, 51, 2415–8.
- Reina, J.; Padilla, E. et al. Evaluation of a new dot blot enzyme immunoassay (directigen flu A+B) for simultaneous and differential detection of influenza A and B virus antigens from respiratory samples. J. Clin.Microbiol. 2002 Sep, 40(9), 3515–17.
Robin T. Clark, Ph.D., is scientific content manager at MilliporeSigma, 28820 Single Oak Dr., Temecula, Calif. 92590, U.S.A.; tel.: 951-326-2755; e-mail: [email protected]; www.emdmillipore.com