Featured Article
Cell free DNA (cfDNA) is present in human blood at varying levels, generally from 1 to 50 ng per mL of plasma. It is typically 100–500 bp in length (with most fragments about 160–180 bp long), making traditional DNA extraction difficult. The key to successful analysis of cfDNA is to ensure efficient purification of smaller DNA fragments (100–500 bp). For this study, the researchers used the cfPure cell free DNA extraction kit from BioChain (Newark, CA), which efficiently recovers these important small fragments. The kit employs a magnetic bead-based DNA extraction method for the recovered cell free DNA. For sensitive applications such as next-generation sequencing, kit performance is comparable to currently available spin column-based cfDNA kits. The recovered DNA is suitable for a wide range of downstream applications, such as bisulfite sequencing, NGS, and qPCR.
Capillary gel electrophoresis system
For this work, the Qsep1 portable bioanalysis system (BiOptic, Inc., New Taipei City, Taiwan [ROC]) was used. Qsep1 utilizes miniature capillary gel electrophoresis (CGE) as the core technology and pen-shaped gel cartridges for separation and fluorescence detection of cfDNA.1
The Qsep1 fragment analyzer is a simple-to-use system that integrates microcapillary electrophoretic technology into a plug-and-play disposable pen-shaped gel cartridge with automated liquid handling and real-time fluorescence detection and analysis. Features include 4–8 sample capacity, a reusable gel cartridge (preprogrammed for 100–200 samples run), and a miniature external dc air pump for a variety of DNA/RNA applications (Figures 1–3). Optimized for high resolution at ambient temperatures, the fragment analyzer provides improved peak resolution, high detection sensitivity, good linear dynamic range, and reproducible migration times, with a reduced instrument and sample analysis cost.
 Figure 1 – Automated detection of cfDNA; sample to result in less than 3 minutes.
Figure 1 – Automated detection of cfDNA; sample to result in less than 3 minutes. Figure 2 – Qsep1 capillary gel electrophoresis system with 4–8 well sample capacity and disposable gel cartridge.
Figure 2 – Qsep1 capillary gel electrophoresis system with 4–8 well sample capacity and disposable gel cartridge.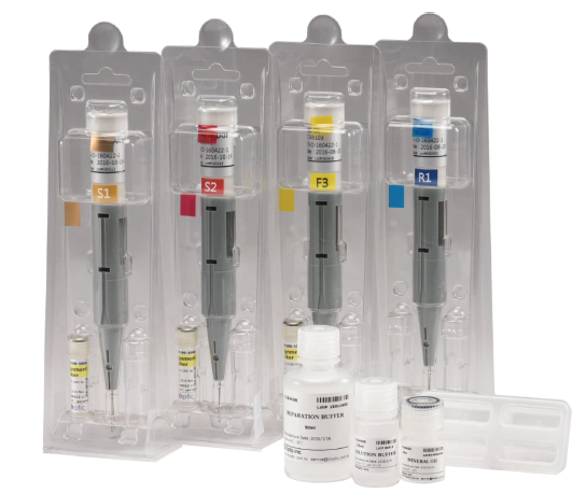 Figure 3 – Reusable pen-shaped gel cartridges.
Figure 3 – Reusable pen-shaped gel cartridges.Results
The cfPure cell free DNA extraction kit was used for the rapid and efficient purification of circulating cfDNA, which was transferred to the Qsep1 system for electrophoretic separation and fluorescence detection (Figures 1 and 2). Silica-coated paramagnetic particles are utilized in the cfPure kit to purify cfDNA from less than 1 mL to greater than 10 mL of serum or plasma. The buffers were developed to ensure efficient recovery of 100–500 bp DNA fragments to maximize recovery of cell free DNA.
cfDNA separation and detection were achieved by using the high-detection-sensitivity N1 gel cartridge (Figure 3). The DNA ladder (pBR322 MspI Digest diluted 100× from the 10 ng/µL stock solution) was used in conjunction with cfDNA samples for accurate size-calling and quantitation.
Electrokinetic injection of the samples was done at 8 kV for 20 seconds, followed by separation at 8 kV for 150 seconds (Figures 4–7). Test results were analyzed using BiOptic Q-Analysis software.
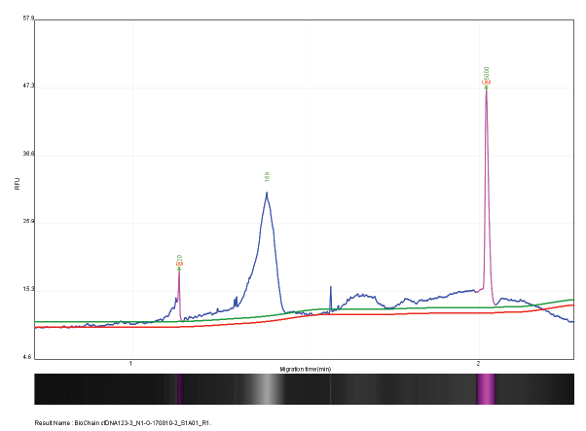 Figure 4 – Cell free DNA (test sample #1) at 169 bp with concentration of 3.12 pg/µL.
Figure 4 – Cell free DNA (test sample #1) at 169 bp with concentration of 3.12 pg/µL.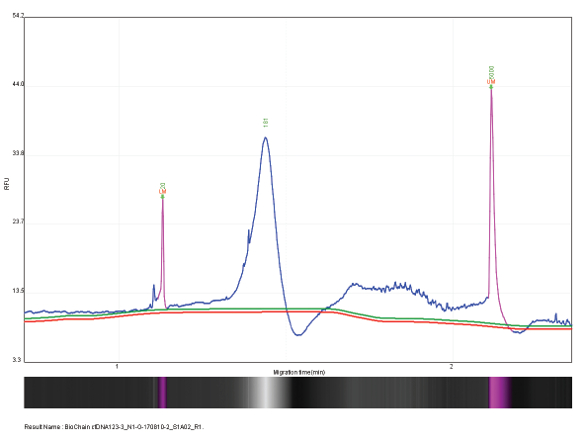 Figure 5 – Cell free DNA (test sample #2) at 181 bp with concentration of 3.82 pg/µL.
Figure 5 – Cell free DNA (test sample #2) at 181 bp with concentration of 3.82 pg/µL. Figure 6 – Cell free DNA (test sample #3) at 171 bp with concentration of 8.21 pg/µL.
Figure 6 – Cell free DNA (test sample #3) at 171 bp with concentration of 8.21 pg/µL.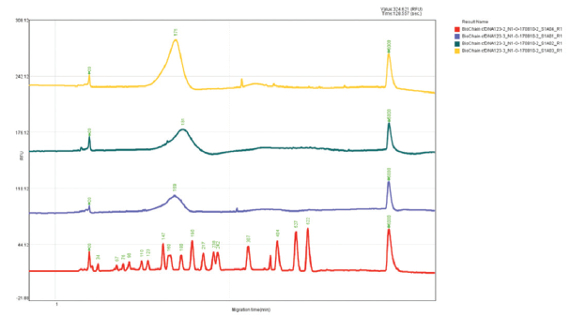 Figure 7 – Cell free DNA test results and DNA ladder (transposed electropherograms).
Figure 7 – Cell free DNA test results and DNA ladder (transposed electropherograms).Conclusion
The portable bioanalysis system described provides high-resolution, high-detection-sensitivity, cost-effective cfDNA analysis for the noninvasive detection of cancer and prenatal genetic testing.
Reference
- www.americanlaboratory.com/914-Application-Notes/339957-A-Field-Portable-Bioanalysis-System/
Varouj Amirkhanian is with BiOptic, Inc., 1409½ Foothill Blvd., La Canada Flintridge, CA 91001, U.S.A.; e-mail:[email protected] or [email protected]; www.bioptic.com.tw. Alice Lin and Eric Tsai are with BiOptic, Inc., New Taipei City, Taiwan (ROC). Geoffrey Routh is with BioChain Institute Inc., Newark, CA.